A graph created by University of Minnesota researchers could lead to more treatment options for COVID-19.
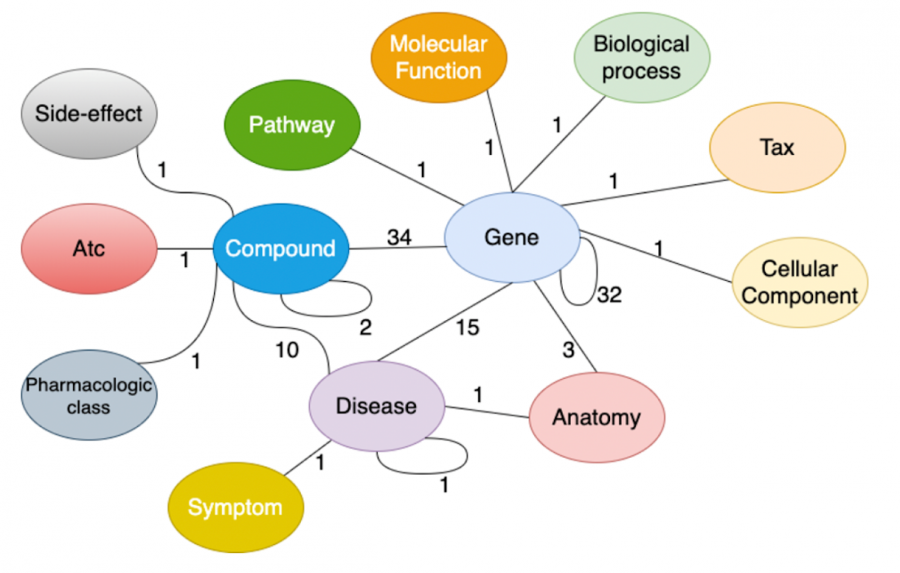
Researchers compiled information from multiple COVID-19 databases to create the Drug Repurposing Knowledge Graph, which shows the relationship between genes, compounds, diseases, biological processes, side effects and symptoms of the disease.
The graph shows how each component is linked and reveals potential interactions. Pre-existing drugs are included to determine whether they may be able to treat other diseases.
University researchers said the goal of the graph is to provide other researchers with a list of drugs that will narrow their search, reduce time and save money.
“The basic idea of drug repurposing is you’re trying to find interactions among, essentially, drugs and diseases. Or, if you want to be a little bit more biological about it, you want to find interactions among drugs and genes that are associated with diseases.” said Vasileios Ioannidis, a researcher and fifth-year University Ph.D. student.
Each disease is associated with a set of genes, he said, with viruses encoded in the genes. Drugs typically relate to the gene in some way, like inhibiting a gene by preventing that gene from expressing another one.
“By following this high-level idea, we can follow this knowledge graph that if a drug is used to inhibit a certain gene and this gene has been found to be related to COVID-19 … then we can have some kind of prediction,” Ioannidis said.
George Karypis, a University professor in the Department of Computer Science and Engineering, has been working on using computational methods like knowledge graphs for solving drug discovery problems for the last 15 years.
By using a knowledge graph to predict missing links and find incomplete information, researchers can discover potential treatments for diseases, he said.
The research group, which includes researchers from the Ohio State University, Hunan University, and Amazon’s AWS AI labs in Shanghai, China and Palo Alto, California, has made the graph publicly available.
“People can just use the knowledge graph and the sample code provided so they can do their own studies,” Karypis said.
The researchers are still working on the knowledge graph, which can be used for diseases beyond COVID-19. In the future, Karypis said they hope to develop a new set of methods that will help people to better leverage the drug knowledge graph for drug repurposing.
“At the end of the day, can we reduce [the] time to essentially ending the pandemic?” he said.