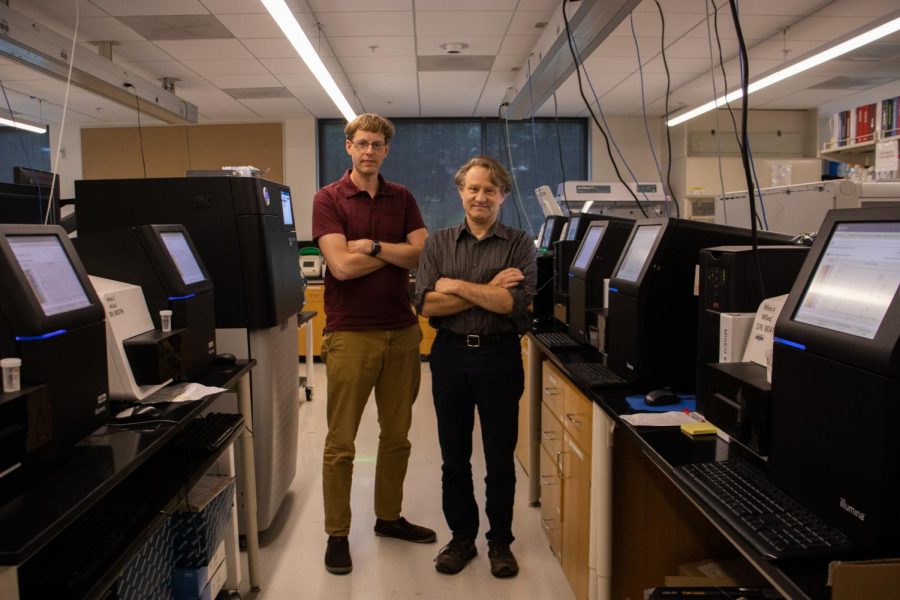
The University of Minnesota Genomics Center (UMGC) recently received funding to conduct genome sampling for COVID-19 in an effort to combat the spread of variants and mutations in the future.
The UMGC, in collaboration with the Minnesota Department of Health (MDH) and with funding from the Center for Disease Control and Prevention (CDC), is working to establish the genetic baseline of the virus in order to prepare for future COVID-19 complications. Although COVID-19 vaccinations are widely available in Minnesota, it is still critical to monitor the viral strains for potential mutations, according to Sean Wang, the sequencing and bioinformatics supervisor at the MDH public health laboratory.
The UMGC developed the sequencing method in May 2020. When people test positive for COVID-19, their tests are sent to labs for genome sequencing, where strands of DNA are studied. Over the next year, the CDC will fund the University to conduct sequencing on 6,000 samples from nine states in the Midwest, including Minnesota, Wang said.
Sequencing genomes is particularly important now in order to prevent the spread of variants, said Kenny Beckman, director of the UMGC.
“Now is the time to treat every single instance as an ember you need to stamp out. If you have a raging forest fire, you just escape the forest fire, but when you have the occasional ember, you can go in there and stamp it out,” Beckman said.
The more people that get infected with COVID-19, the more variants are able to emerge, as each new host gives the virus another chance to mutate and adapt, according to Daryl Gohl, the principal investigator from the University and genomics professor.
“I think there will still be work to do to investigate the outbreaks,” Gohl said. “It is very important to keep tabs on that and to make sure that there aren’t variants arising that are becoming resistant or evading the immune system of vaccinated people.”
The low number of COVID-19 outbreaks in Minnesota has made it much easier to sequence all of the samples, Beckman said. When there were high case numbers, it was not possible to sequence each test. MDH collaborates with the UMGC to help determine which samples to sequence. In some cases, samples from an outbreak investigation are sequenced, and other times random samples are sequenced for generalized surveillance.
“Our job is still critical to closely monitor whether or not we have a surge of the [most contagious] variant. Or, who knows what kind of variant we’ll have once we step closer into the fall/winter time,” Wang said. “The term restrictions have loosened up so there are more opportunities for the virus to spread. So we definitely will not put our guards down.”
Correction: A previous version of this story included incorrect wording of a quote from Daryl Gohl.
























Rusty
Oct 14, 2021 at 12:17 pm
To figure out how to kill off 90% of the world population, to “save” the planet.